Muscle fiber segmentation software implemented as an ImageJ macro.
DESCRIPTION
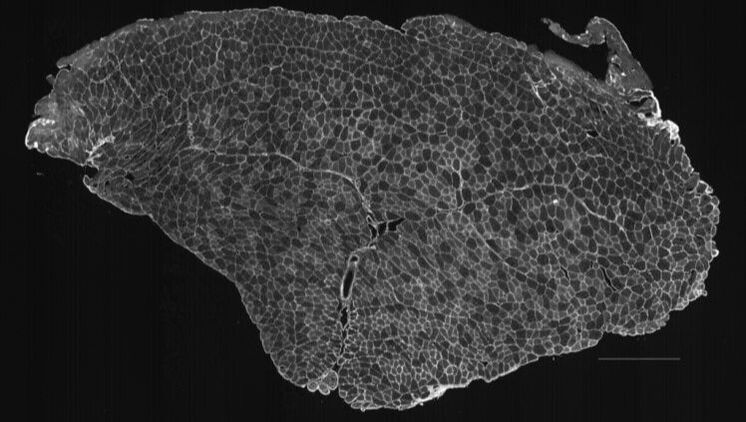
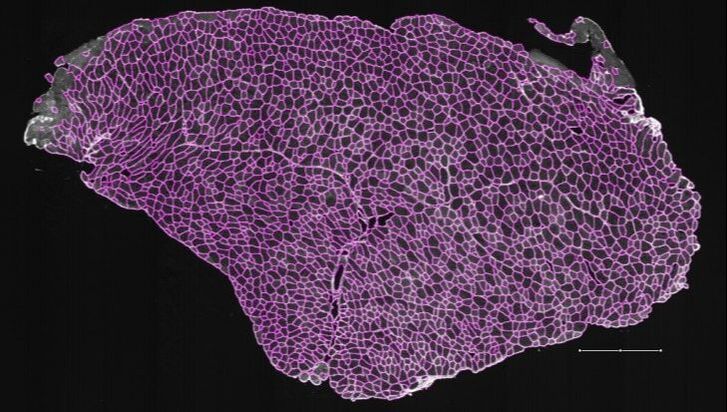
MyoSAT is an ImageJ Macro application developed for automated segmentation of muscle cross-section images for histology analysis. The input is an image of a muscle cross-section. The image processing routines automatically segment the muscle fibers in the image. Results include an overlay image of the segmentation result as well as fiber size histograms and count data.
Input images must be either 8 bit or 16 bit grayscale *.tiff format. (tiled format tiffs generated by slide scanners must first be converted to flat format – see user manual)
MyoSAT was originally developed to analyze skeletal muscle labeled with Collagen V antibody using fluorescence immunohistochemistry to label the fiber borders. The software will work with other staining methods including H&E staining if the input image is properly formatted.
Our research group developed MyoSAT after having limited success analyzing large muscle cross-section scans using available software. The image processing pipeline has been optimized to work in challenging cases of weak or uneven staining contrast of the muscle. The macro has been tested to allow automatic segmentation of large area (whole slide) muscle cross-sections containing thousands of fibers - provided sufficient computer memory is available. The software is easy to configure and use.
MyoSAT was developed by the Cheetham Research Group at the Cornell University Veterinary College.
MyoSAT is released as an open source project for use and continued development by the research community.
MyoSAT is an ImageJ Macro application developed for automated segmentation of muscle cross-section images for histology analysis. The input is an image of a muscle cross-section. The image processing routines automatically segment the muscle fibers in the image. Results include an overlay image of the segmentation result as well as fiber size histograms and count data.
Input images must be either 8 bit or 16 bit grayscale *.tiff format. (tiled format tiffs generated by slide scanners must first be converted to flat format – see user manual)
MyoSAT was originally developed to analyze skeletal muscle labeled with Collagen V antibody using fluorescence immunohistochemistry to label the fiber borders. The software will work with other staining methods including H&E staining if the input image is properly formatted.
Our research group developed MyoSAT after having limited success analyzing large muscle cross-section scans using available software. The image processing pipeline has been optimized to work in challenging cases of weak or uneven staining contrast of the muscle. The macro has been tested to allow automatic segmentation of large area (whole slide) muscle cross-sections containing thousands of fibers - provided sufficient computer memory is available. The software is easy to configure and use.
MyoSAT was developed by the Cheetham Research Group at the Cornell University Veterinary College.
MyoSAT is released as an open source project for use and continued development by the research community.
IMAGE PROCESSING APPROACH
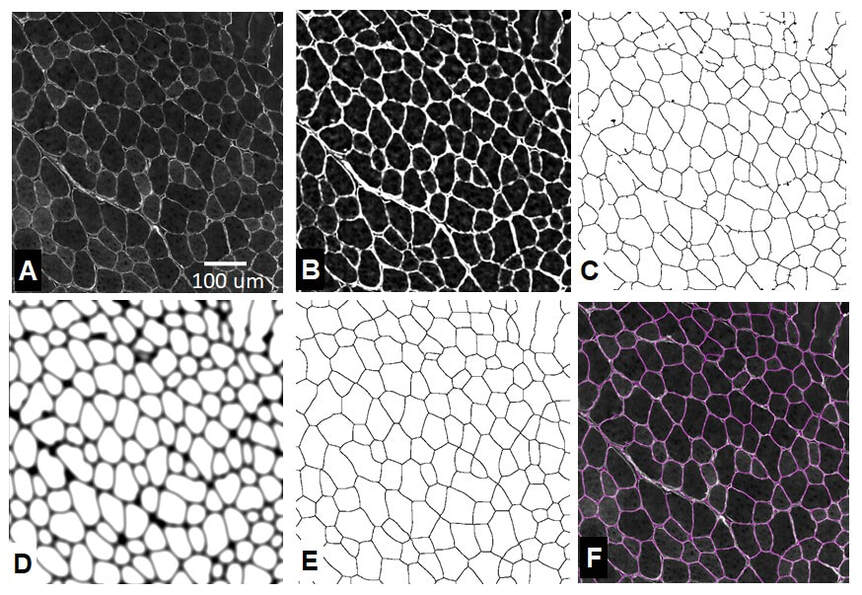
The image processing approach involves three main stages: The first stage is pre-processing. The image is background leveled to compensate for uneven staining intensity. This is followed by contrast enhancement steps. In the second stage, the macro makes use of Steger’s line detection algorithm to locate fiber boundary candidates. After additional processing steps, The third stage employs classical watershed segmentation which is used to generate the final segmentation. Fiber sizes are reported by minimum feret diameter as well as other metrics. Details of the algorithm development are detailed in the research paper below:
Development of the processing pipeline is described in this paper:
Approach for Semi-Automated Measurement of Fiber Diameter in Murine and Canine Skeletal Muscle.
Authors: Courtney R. Stevens, Michael Sledziona, Josh Berenson, Timothy P. Moore, Lynn Dong, Jonathan Cheetham
Preprint submitted 3/7/2019 [PREPRINT: https://www.biorxiv.org/content/10.1101/569780v1]
Briefly: The initial image processing starts with leveling steps to adjust for uneven staining followed by contrast enhancements. The leveling is done by selectively enhancing image regions to match a uniform median background level. The macro employs the use of Steger’s line detection algorithm to locate the fiber boundary candidates. After additional processing steps, classical watershed segmentation is used to generate the final segmentation. Fiber sizes are reported by minimum feret diameter.
The image processing approach involves three main stages: The first stage is pre-processing. The image is background leveled to compensate for uneven staining intensity. This is followed by contrast enhancement steps. In the second stage, the macro makes use of Steger’s line detection algorithm to locate fiber boundary candidates. After additional processing steps, The third stage employs classical watershed segmentation which is used to generate the final segmentation. Fiber sizes are reported by minimum feret diameter as well as other metrics. Details of the algorithm development are detailed in the research paper below:
Development of the processing pipeline is described in this paper:
Approach for Semi-Automated Measurement of Fiber Diameter in Murine and Canine Skeletal Muscle.
Authors: Courtney R. Stevens, Michael Sledziona, Josh Berenson, Timothy P. Moore, Lynn Dong, Jonathan Cheetham
Preprint submitted 3/7/2019 [PREPRINT: https://www.biorxiv.org/content/10.1101/569780v1]
Briefly: The initial image processing starts with leveling steps to adjust for uneven staining followed by contrast enhancements. The leveling is done by selectively enhancing image regions to match a uniform median background level. The macro employs the use of Steger’s line detection algorithm to locate the fiber boundary candidates. After additional processing steps, classical watershed segmentation is used to generate the final segmentation. Fiber sizes are reported by minimum feret diameter.
REFERENCES
Approach for Semi-Automated Measurement of Fiber Diameter in Murine and Canine Skeletal Muscle.
Authors: Courtney R. Stevens, Michael Sledziona, Josh Berenson, Timothy P. Moore, Lynn Dong, Jonathan Cheetham
Preprint submitted 3/7/2019 [PREPRINT: https://www.biorxiv.org/content/10.1101/569780v1]
C. Steger, "An unbiased detector of curvilinear structures," in IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 20, no. 2, pp. 113-125, Feb. 1998, doi: 10.1109/34.659930
IJ-Plugin Toolkit: http://ij-plugins.sourceforge.net/plugins/toolkit.html
Legland, D.; Arganda-Carreras, I. & Andrey, P. (2016), "MorphoLibJ: integrated library and plugins for mathematical morphology with ImageJ", Bioinformatics (Oxford Univ Press) 32(22): 3532-3534, PMID 27412086, doi:10.1093/bioinformatics/btw413
Steger’s Ridge Detection Inplementation: thorstenwagner/ij-ridgedetection: Ridge Detection 1.4.0 https://zenodo.org/record/845874#.XrVo3ahKguU
Approach for Semi-Automated Measurement of Fiber Diameter in Murine and Canine Skeletal Muscle.
Authors: Courtney R. Stevens, Michael Sledziona, Josh Berenson, Timothy P. Moore, Lynn Dong, Jonathan Cheetham
Preprint submitted 3/7/2019 [PREPRINT: https://www.biorxiv.org/content/10.1101/569780v1]
C. Steger, "An unbiased detector of curvilinear structures," in IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 20, no. 2, pp. 113-125, Feb. 1998, doi: 10.1109/34.659930
IJ-Plugin Toolkit: http://ij-plugins.sourceforge.net/plugins/toolkit.html
Legland, D.; Arganda-Carreras, I. & Andrey, P. (2016), "MorphoLibJ: integrated library and plugins for mathematical morphology with ImageJ", Bioinformatics (Oxford Univ Press) 32(22): 3532-3534, PMID 27412086, doi:10.1093/bioinformatics/btw413
Steger’s Ridge Detection Inplementation: thorstenwagner/ij-ridgedetection: Ridge Detection 1.4.0 https://zenodo.org/record/845874#.XrVo3ahKguU
Software Copyright Notice: Copyright (c)2020 Cornell University
For Public Release
Authors: Courtney Stevens, Michael Sledziona, Josh Berenson (Cornell University College of Veterinary Medicine)
Primary Contacts:
Michael Sledziona ms66 {at} cornell.edu
Dr. Jonathan Cheetham jc485 {at} cornell.edu
For Public Release
Authors: Courtney Stevens, Michael Sledziona, Josh Berenson (Cornell University College of Veterinary Medicine)
Primary Contacts:
Michael Sledziona ms66 {at} cornell.edu
Dr. Jonathan Cheetham jc485 {at} cornell.edu
|
© 2020 Jon Cheetham Lab | Cornell University |